Compiled Extensions: The Shared Object Pipeline
Last updated on 2026-02-12 | Edit this page
Estimated time: 50 minutes
Overview
Questions
- How does the Python ecosystem deliver high-performance binaries to users?
- What distinguishes a “Source Distribution” from a “Binary Wheel”?
- What role does
cibuildwheelplay in software distribution?
Objectives
- Construct a Python extension module from a Fortran kernel using Meson.
- Configure a build-backend for compiled extensions using
meson-python. - Setup a CI pipeline to generate binary wheels for distribution.
The Shared Object Reality
In the domain of scientific Python, “packaging” often refers
effectively to “binary distribution.” When users install libraries such
as numpy, torch, or openblas, they typically download compiled
artifacts rather than pure Python scripts.
Investigation of the site-packages directory reveals
that the core logic resides in Shared Object files
(.so on Linux, .dylib on macOS,
.dll on Windows). Python functions primarily as the
interface.
To distribute high-performance code effectively, one must master the pipeline that generates these artifacts consisting of
- Translation
- Generating C wrappers for Fortran/C++ code.
- Compilation
- Transforming source code into shared objects.
- Bundling
-
Packaging shared objects into Wheels (
.whl).
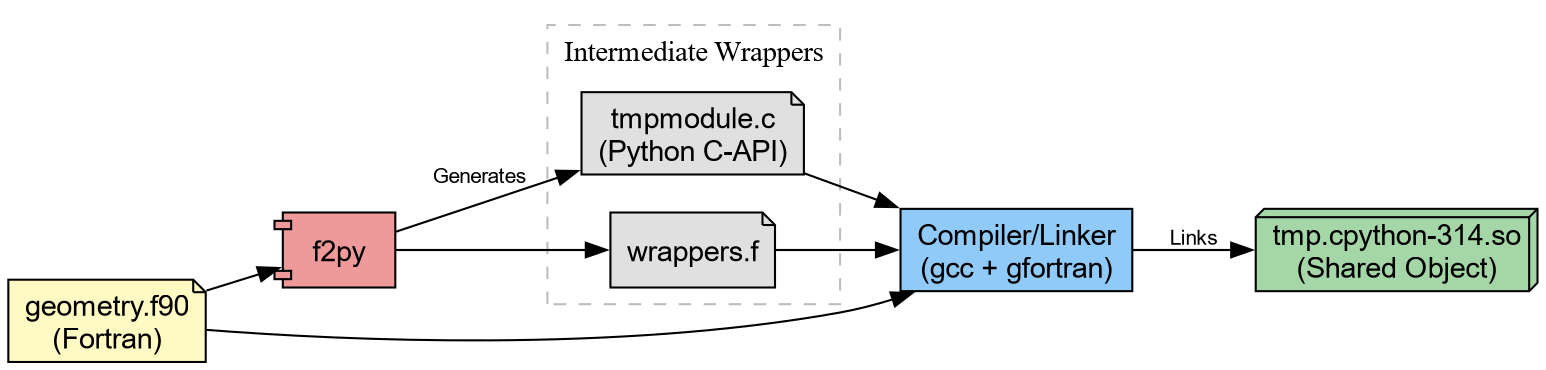
The Computational Kernel in Fortran
We begin with a computational kernel. In physical chemistry, calculating the Euclidean distance between atomic coordinates constitutes a fundamental operation.
Create src/chemlib/geometry.f90:
F90
subroutine calc_distance(n, r_a, r_b, dist)
implicit none
integer, intent(in) :: n
real(8), intent(in), dimension(n) :: r_a, r_b
real(8), intent(out) :: dist
!f2py intent(hide) :: n
!f2py intent(in) :: r_a, r_b
!f2py intent(out) :: dist
integer :: i
dist = 0.0d0
do i = 1, n
dist = dist + (r_a(i)-r_b(i))**2
end do
dist = sqrt(dist)
end subroutine calc_distanceThis can be immediately compiled through f2py.
Which generates tmp.cpython*.so (or .dll on
Windows). We can try this out.
PYTHON
❯ python
Python 3.14.2 (main, Jan 2 2026, 14:27:39) [GCC 15.2.1 20251112] on linux
Type "help", "copyright", "credits" or "license" for more information.
>>> import tmp
>>> tmp.calc_distance([0.,0.,0.], [1.,1.,1.])
1.7320508075688772Challenge: Validate the result
Can you make sure the results are correct? Note that since we cannot
really “install” the package yet, a simple assert will have to do.
Considering “installable” variants with meson
The problem with the setup so far is that the
compiled extension doesn’t get “installed” into site-packages. To work around this, it helps to
first take a look behind the curtain. f2py with the
--build-dir option will output the intermediate build
files.
BASH
❯ f2py -c geometry.f90 -m tmp --build-dir tmp_out
Cannot use distutils backend with Python>=3.12, using meson backend instead.
Using meson backend
Will pass --lower to f2py
See https://numpy.org/doc/stable/f2py/buildtools/meson.html
Reading fortran codes...
Reading file 'geometry.f90' (format:free)
Post-processing...
character_backward_compatibility_hook
Post-processing (stage 2)...
Building modules...
Building module "tmp"...
Generating possibly empty wrappers"
Maybe empty "tmp-f2pywrappers.f"
Constructing wrapper function "calc_distance"...
dist = calc_distance(r_a,r_b)
Wrote C/API module "tmp" to file "./tmpmodule.c"
The Meson build system
Version: 1.10.1
Source dir: /home/rgoswami/Git/Github/epfl/pixi_envs/teaching/python_packaging_workbench/python_packaging_workbench/org_src/episodes/data/chemlib/src/chemlib/tmp_out
Build dir: /home/rgoswami/Git/Github/epfl/pixi_envs/teaching/python_packaging_workbench/python_packaging_workbench/org_src/episodes/data/chemlib/src/chemlib/tmp_out/bbdir
Build type: native build
Project name: tmp
Project version: 0.1
Fortran compiler for the host machine: gfortran (gcc 15.2.1 "GNU Fortran (GCC) 15.2.1 20260103")
Fortran linker for the host machine: gfortran ld.bfd 2.45.1
C compiler for the host machine: cc (gcc 15.2.1 "cc (GCC) 15.2.1 20260103")
C linker for the host machine: cc ld.bfd 2.45.1
Host machine cpu family: x86_64
Host machine cpu: x86_64
Program /usr/bin/python found: YES (/usr/bin/python)
Found pkg-config: YES (/usr/bin/pkg-config) 2.5.1
Build targets in project: 1
Found ninja-1.13.2 at /usr/bin/ninja
INFO: autodetecting backend as ninja
INFO: calculating backend command to run: /usr/bin/ninja -C /home/rgoswami/Git/Github/epfl/pixi_envs/teaching/python_packaging_workbench/python_packaging_workbench/org_src/episodes/data/chemlib/src/chemlib/tmp_out/bbdir
ninja: Entering directory `/home/rgoswami/Git/Github/epfl/pixi_envs/teaching/python_packaging_workbench/python_packaging_workbench/org_src/episodes/data/chemlib/src/chemlib/tmp_out/bbdir'
[7/7] Linking target tmp.cpython-314-x86_64-linux-gnu.soWhich we can then inspect..
The intermediate involves a meson.build !
project('tmp',
['c', 'fortran'],
version : '0.1',
meson_version: '>= 1.1.0',
default_options : [
'warning_level=1',
'buildtype=release'
])
fc = meson.get_compiler('fortran')
py = import('python').find_installation('''/usr/bin/python''', pure: false)
py_dep = py.dependency()
incdir_numpy = run_command(py,
['-c', 'import os; os.chdir(".."); import numpy; print(numpy.get_include())'],
check : true
).stdout().strip()
incdir_f2py = run_command(py,
['-c', 'import os; os.chdir(".."); import numpy.f2py; print(numpy.f2py.get_include())'],
check : true
).stdout().strip()
inc_np = include_directories(incdir_numpy)
np_dep = declare_dependency(include_directories: inc_np)
incdir_f2py = incdir_numpy / '..' / '..' / 'f2py' / 'src'
inc_f2py = include_directories(incdir_f2py)
fortranobject_c = incdir_f2py / 'fortranobject.c'
inc_np = include_directories(incdir_numpy, incdir_f2py)
# gh-25000
quadmath_dep = fc.find_library('quadmath', required: false)
py.extension_module('tmp',
[
'''geometry.f90''',
'''tmpmodule.c''',
'''tmp-f2pywrappers.f''',
fortranobject_c
],
include_directories: [
inc_np,
],
objects: [
],
dependencies : [
py_dep,
quadmath_dep,
],
install : true)We could take inspiration from this, generate sources, and link them together:
f2py_prog = find_program('f2py')
# Generate Wrappers
geometry_source = custom_target('geometrymodule.c',
input : ['src/chemlib/geometry.f90'],
output : ['geometrymodule.c', 'geometry-f2pywrappers.f'],
command : [f2py_prog, '-m', 'geometry', '@INPUT@', '--build-dir', '@OUTDIR@']
)A pattern commonly used in SciPy for instance. Here we will consider
a less tool-heavy approach though build backends such as meson-python, scikit-build-core, and setuptools orchestrate complex builds.
The “Manual” Install
For now, let’s consider falling back to what we learned about installation.
Challenge: The Site-Packages Hack
Your goal: Make the tmp module importable from anywhere
in your system (within the current environment), not just the source
folder.
Locate the active site-packages directory for your
current environment.
Copy the compiled .so (or .pyd) file into
that directory.
Change your directory to $HOME (to ensure you do not
import the local file).
Launch Python and attempt to import tmp.
This manual exercise mimics exactly how libraries like openblas, metatensor, and torch operate. If you examine their installed
folders, you will find large compiled shared objects.
The Python files (__init__.py) serve
mostly as wrappers to load these binary blobs. For example, a robust
package might look like this:
PYTHON
try:
from . import _geometry_backend
except ImportError: # Logic to handle missing binaries or wrong platforms
raise ImportError("Could not load the compiled extension!")
def calc_distance(a, b): # Pure Python type checking before passing to Fortran
return _geometry_backend.calc_distance(a, b)The Wheelhouse
We now possess a working shared object. However, a critical flaw remains: Portability.
The .so file you just generated links against:
- The specific version of Python on your machine.
- The system C library (glibc) on your machine.
- The CPU architecture (x8664, ARM64) of your machine.
If you email this file to a colleague running Windows, or even a different version of Linux, it will crash.
To distribute this code, we cannot ask every user to install a
Fortran compiler and run f2py. Instead, we use
cibuildwheel to distribute binaries to end users.
A “Wheel” (.whl) functions as a ZIP archive containing
the artifacts we just manually moved. To support the community, we must
generate wheels for every combination of:
- Operating System (Windows, macOS, Linux)
- Python Version (3.10, 3.11, 3.12, 3.13, 3.14)
- Architecture (x86, ARM)
Tools like cibuildwheel automate this matrix. They spin
up isolated environments (often using Docker or virtual machines),
compile the code, fix the library linkages (bundling dependencies), and
produce the final artifacts.
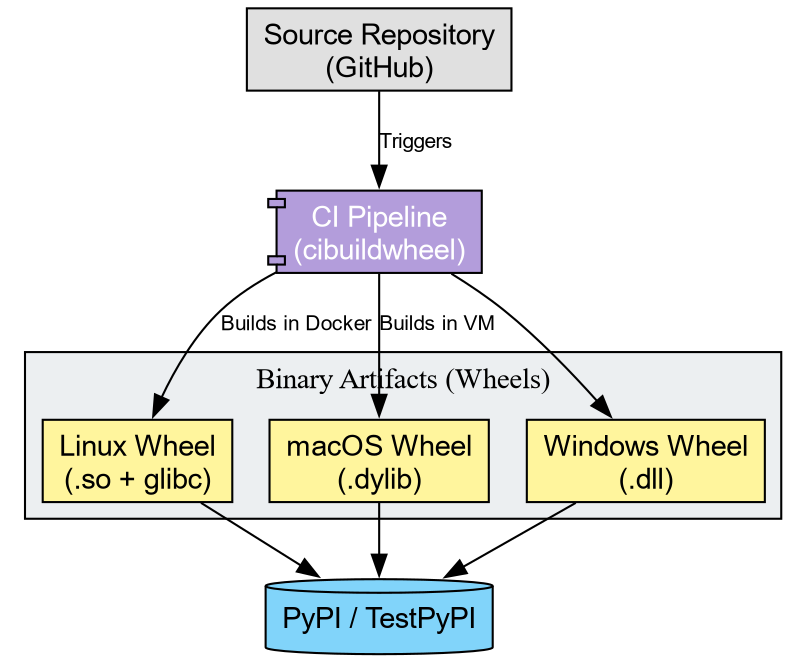
Challenge: Conceptualizing the Pipeline
Imagine you publish chemlib. A user reports:
“ImportError: DLL load failed: The specified module could not be found.”
Based on today’s lesson, what likely went wrong?
- The Python code has a syntax error.
- The user’s computer lacks a Fortran compiler.
- The specific shared object for their OS/Python version was missing or incompatible.
Answer: 3.
The error “DLL load failed” implies the Python interpreter attempted to load the shared object but failed. This usually occurs when the binary wheel does not match the user’s system, or the wheel failed to bundle a required system library. The user does not need a compiler (Option 2) if they are using a Wheel.
The Manylinux Standard
On Linux, binary compatibility presents a challenge due to varying
system libraries (glibc). cibuildwheel addresses this by
executing the build inside a specialized Docker container (Manylinux).
This ensures the compiled .so file links against an older
version of glibc, guaranteeing functionality on the majority of Linux
distributions.
Challenge: Inspecting an Artifact
- Go to the PyPI page for
metatomic. - You will observe a file ending in
.whl. - Treat this file as a ZIP archive (which it represents). Unzip it.
- Locate the
.so(or.pyd) file inside.
Reflection: This binary file constitutes the actual product consumed by users. The Fortran source code effectively disappears, becoming baked into the machine code of this shared object.
-
Shared Objects: Scientific Python packages function
primarily as delivery mechanisms for compiled binaries (
.sofiles). - Installation: “Installing” a package physically amounts to copying these binaries into site-packages.
- Cibuildwheel: Automates the creation of binary wheels for all platforms, removing the need for users to possess compilers.
